Biologia plantarum 70:27-37, 2026 | DOI: 10.32615/bp.2026.004
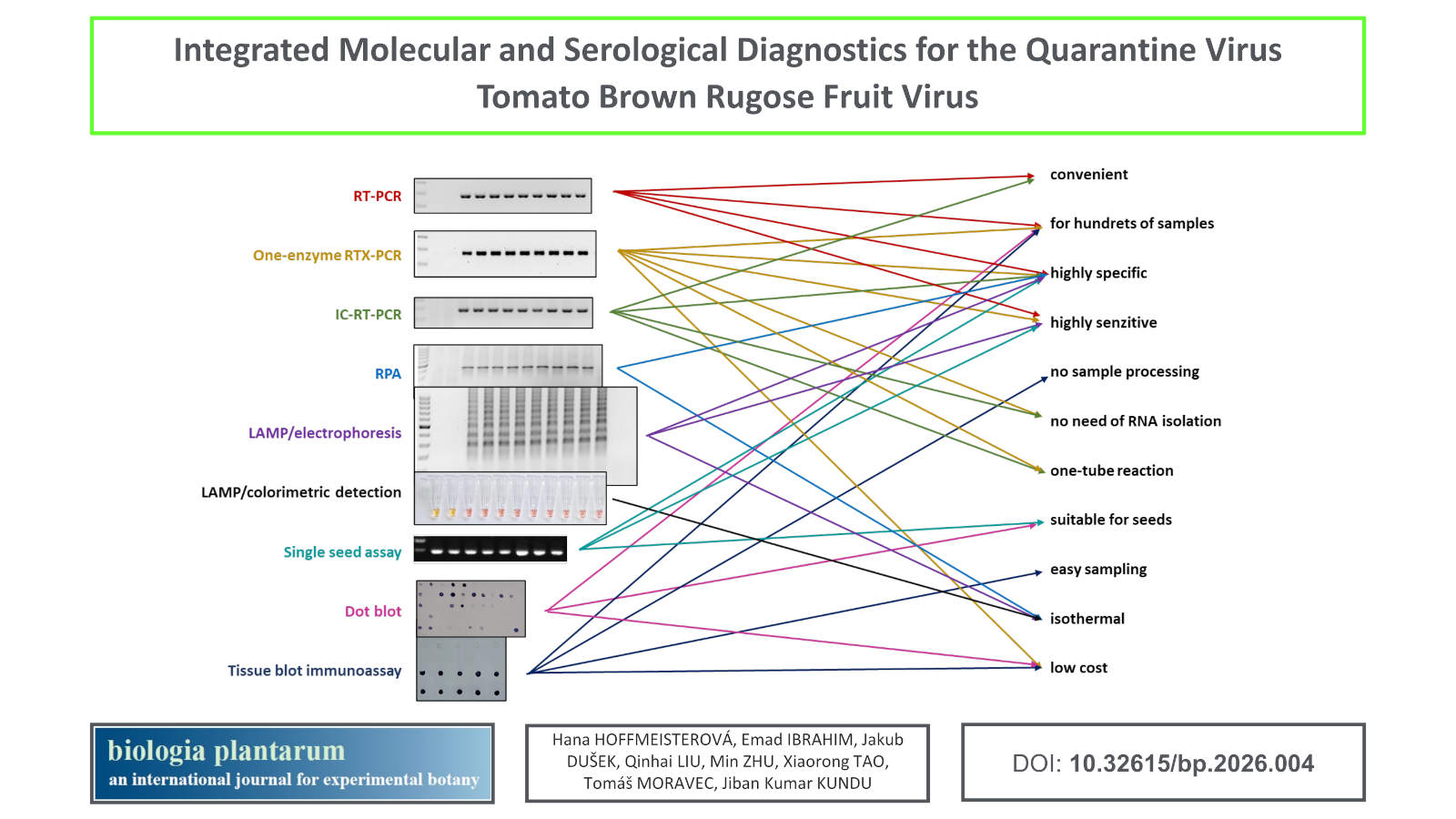
Integrated molecular and serological diagnostics for surveillance of the quarantine virus tomato brown rugose fruit virus
- 1 Plant Virus and Vector Interactions, Centre for Plant Virus Research, Czech Agrifood Research Center, Drnovská 507, 161 06 Prague, Czech Republic
- 2 Laboratory of Virology, Centre for Plant Virus Research, Institute of Experimental Botany of the Czech Academy of Sciences, Rozvojová 263, 165 00 Prague, Czech Republic
- 3 Department of Plant Pathology, Nanjing Agricultural University, 210095 Nanjing, People's Republic of China
Background: The tomato brown rugose fruit virus (Tobamovirus fructirugosum, ToBRFV) is an emerging tobamovirus that has quickly become a significant obstacle to the production of tomatoes and peppers worldwide. It is now classified as a regulated quarantine pathogen. Effective containment requires rapid, reliable, inexpensive, and safe diagnostic protocols for routine screening in laboratories and production systems.
Aims: We aimed to thoroughly evaluate integrated molecular and serological diagnostic methods for ToBRFV and develop biosafe positive controls suitable for high-throughput and decentralized applications.
Methods: We evaluated the following methods: conventional RT-PCR, one-enzyme RTX-PCR, immunocapture RT-PCR, recombinase polymerase amplification, loop-mediated isothermal amplification with colorimetric detection, Western blotting, dot blot, and tissue blot immunoassay. The non-infectious positive control was prepared using the GoldenBraid 3.0 cloning system. We developed a single-seed assay that enables direct testing of tomato seed stocks.
Results: Among the evaluated molecular methods, RTX-PCR was particularly advantageous due to its minimal sample handling, reduced cost, and ability to bypass RNA extraction. The tissue blot immunoassay enabled high-throughput, low-cost screening of hundreds of samples per day using only basic equipment. Although ToBRFV was frequently detected in seeds harvested from infected plants, no systemic infection was observed in progeny seedlings, confirming the low rate of true vertical transmission. A non-infectious positive control was prepared and successfully employed in molecular methods.
Conclusions: Our findings provide an integrated diagnostic framework combining molecular, serological, and biosafety tools to effectively monitor and contain ToBRFV in commercial production and phytosanitary settings.
Keywords: colorimetric LAMP detection, one-enzyme RTX PCR, quarantine plant virus, tissue blot immunoassay (TBIA), tomato brown rugose fruit virus.
Received: December 11, 2025; Revised: February 27, 2026; Accepted: March 5, 2026; Published online: March 18, 2026 Show citation
| ACS | AIP | APA | ASA | Harvard | Chicago | Chicago Notes | IEEE | ISO690 | MLA | NLM | Turabian | Vancouver |
Introduction
Tomato brown rugose fruit virus (Tobamovirus fructirugosum, ToBRFV), a member of the genus Tobamovirus, has emerged over the past decade as one of the most damaging viral pathogens of tomato (Solanum lycopersicum) and pepper (Capsicum spp.) worldwide (Caruso et al., 2022). First identified in greenhouse grown tomatoes in Jordan (Salem et al., 2016), ToBRFV is characterised by its high contagiosity and its ability to overcome the Tm-22 resistance gene, which has long been a cornerstone of tobamovirus resistance breeding in tomato (Luria et al., 2018; Maayan et al., 2018). Its efficient mechanical and seed-associated transmission, together with its remarkable environmental stability and absence of effective control measures, have led according to the European and Mediterranean Plant Protection Organization (EPPO) to its recent designation as a quarantine pathogen in Morocco (since 2020) and China (since 2021) and A1 or A2 list-pathogen in several countries all around the world (EPPO, 2020a).
Since its initial discovery, according to the most recent EPPO report, ToBRFV has spread rapidly on all continents except Antarctica, placing increasing pressure on phytosanitary authorities and diagnostic laboratories. Limiting further spread requires early, sensitive, and rapid diagnostic systems that can keep pace with the virus’s transmission dynamics and support timely phytosanitary decision-making (Islam et al., 2019; Davino et al., 2020).
Early and reliable detection is essential for containing outbreaks, safeguarding international seed and fruit trade, and mitigating severe economic losses. In response, several molecular and serological diagnostic tools have been developed, including RT-qPCR (Menzel and Winter, 2021), loop-mediated isothermal amplification (LAMP) (Sarkes et al., 2020), and tissue blot immunoassay (TBIA) (Dombrovsky and Smith, 2017). These methods offer high sensitivity and versatility, and several are amenable to field deployment, enabling real-time support for quarantine interventions.
A major challenge in diagnostics for regulated viruses such as ToBRFV is the requirement for infectious viral material as positive controls. Handling live virus poses biosafety risks, especially in laboratories without containment facilities or in regions free of ToBRFV. Accidental release during diagnostic workflows could lead to new outbreaks or compromise quarantine integrity (López-Moya et al., 2000). International shipment of infectious virus is also heavily restricted, limiting inter laboratory standardisation and consistent assay validation. Consequently, the development of biosafe positive controls – such as synthetic RNA standards or recombinant plasmids – has become a priority (López-Moya et al., 2000). These controls are safe to store, transport, and use, while remaining fully compatible with molecular assays including RT-qPCR, endpoint RT-PCR, LAMP, and others.
In this study, we provide an integrated evaluation of efficient and environmentally safe diagnostic approaches for ToBRFV. We assess the performance of molecular assays – including endpoint RT-PCR using RTX polymerase (RTX-PCR), RT-LAMP, and recombinase polymerase amplification (RPA) – alongside serological methods such as TBIA. We also describe the development and validation of non-infectious positive controls to enhance biosafety and standardisation. In addition, we establish a single-seed assay enabling direct detection of ToBRFV in individual tomato seeds, a key requirement for quarantine-compliant seed testing.
Collectively, the diagnostic tools evaluated in this study provide sensitive, rapid, and practical alternatives for monitoring ToBRFV in research, regulatory, and production settings, supporting more robust surveillance and effective containment of this economically important quarantine virus.
Materials and methods
Virus source: ToBRFV isolate, originally obtained from Solanum lycopersicum (tomato) in North Rhine Westphalia, Germany, was purchased from the DSMZ – German Collection of Microorganisms and Cell Cultures (Leibniz Institute, Science Campus Braunschweig-Süd; catalogue number PC-1236; GenBank: MK133093).
Plant material: Nicotiana benthamiana plants were used for the initial propagation of ToBRFV and for preparing virus-rich tissue for the virus isolation. For subsequent evaluations of diagnostic methods, tomato plants (Solanum lycopersicum cv. Gallant) served as the primary experimental host. N. benthamiana and tomato cv. Gallant plants were grown in a controlled-environment chamber at 20°C with a 16 h light/8 h dark. To assess ToBRFV infection under production conditions, we collaborated with Hanácká zemědělská společnost Jevíčko a.s. (HZSJ), a commercial tomato producer in the Moravian region of the Czech Republic. This collaboration focused on two tomato cultivars, Plumola and Lucioso, cultivated hydroponically in greenhouses in Chornice during 2023 - 2025. Sampling was done randomly in a general screening. To monitor the potential incidence of ToBRFV, we also analysed samples from a broad range of tomato cultivars (Balkonové, Bejbino, Blumko, Dafne, Delikano, Denár, Gallant, Hroznové, Jergus, Mandat, Merrygold, Nagina, Odat, Orbit, Pálava, Pedro, Perun, Rose Crush, Rose Lady, Rubinka, Start, Tipo, Tornádo, Toro, Valdo) collected randomly in a general screening from private gardens and experimental field plots across the Czech Republic between 2023 and 2025. Tomato cv. Mingzhi88 was used specifically for the development of the single-seed detection assay.
Mechanical inoculation: Lyophilized, commercially purchased leaves containing ToBRFV were rehydrated for 1 h in 10 mM potassium phosphate buffer (pH 7.0) at a (mass:volume) ratio, and subsequently homogenized using a mortar and pestle. The resulting sap was mechanically inoculated onto two leaves per N. benthamiana plant according to Hoffmeisterová et al. (2025). The inoculum for tomato inoculation was prepared from ToBRFV-infected N. benthamiana leaves, which were homogenised in 10 mM potassium phosphate buffer (pH 7.0) at a 1:10 (w/v) ratio using a mortar and pestle. Tomato plants at the four-true-leaf stage were mechanically inoculated as described above.
ToBRFV propagation: To propagate ToBRFV, N. benthamiana plants were used. Approximately 12 days post inoculation (dpi), symptom development was assessed, and ToBRFV infection was confirmed by one-enzyme RTX-PCR (Hoffmeisterová et al., 2022).
ToBRFV isolation: ToBRFV was isolated from systemic N. benthamiana leaves at 14 dpi using a tobacco mosaic virus (Tobamovirus tabaci, TMV) isolation protocol described by Asselin and Zaitlin (1978). Finally, the virus concentration was estimated by UV absorbance using a NanoDrop spectrophotometer (Thermo Scientific, Wilmington, DE, USA), applying the Beer–Lambert law and the molar extinction coefficient for TMV (ε₂₆₀ = 3.0) (Zaitlin, 1975). Purity of the preparation was evaluated by SDS-PAGE followed by Coomassie Brilliant Blue R-250 staining.
Vertical transmission of ToBRFV: Tomato plants confirmed by RTX-PCR to be ToBRFV-positive were transferred to larger pots and maintained until fruit development. Seeds were extracted from ripe fruits, fermented overnight in water at room temperature with continuous stirring, and then cleaned of residual pulp before being air-dried for approximately 1 week.
A first batch of 43 seeds, collected from five fruits (7, 10, 8, 5, and 13 seeds per fruit), was tested individually for ToBRFV. A second batch of 47 seeds was sown and grown under standard greenhouse conditions for 4 weeks. Individual seedlings were subsequently analysed using tissue blot immunoassay (TBIA) and one-enzyme RTX PCR to assess systemic infection by ToBRFV.
In paralell, the tomato single-seed ToBRFV detection system was developed (see below).
Preparation of a safe positive control: A biosafe positive control, designated ToBRFV-pDB1α1/PC-ToBRFV/PVX (previously labelled pDB1a1:PC-ToBRFV-MP/PVX-Sap; Utility Model CZ 37695), was constructed by inserting a fragment of the ToBRFV movement protein (MP) gene into a non-quarantine potato virus X (Potexvirus solani, PVX)-based vector. A 462 bp fragment of the MP gene was amplified by PCR using primers GB-PC-ToBRFV-F and GB-PC-ToBRFV-R (Table 1). These primers contained BsmBI restriction sites (CGTCTC) and four-nucleotide overhangs compatible with the GoldenBraid domestication plasmid pUPD2 (Sarrion-Perdigones et al., 2011; Dusek et al., 2020). Additional four-nucleotide sequences (AATG in the forward primer; GCTT in the reverse primer) facilitated subsequent cloning from pUPD2 into the infectious PVX clone.
PCR reactions contained 100 ng ToBRFV cDNA, 0.4 U Phusion™ High-Fidelity DNA Polymerase, 1× HF buffer (Thermo Fisher Scientific, Waltham, MA, USA), 10 mM each dNTP, 300 nM of each primer and H2O up to 20 µL. The thermal profile consisted of 98°C for 30 s; 35 cycles of 98°C for 10 s, 58°C for 20 s, and 72°C for 30 s; followed by a final extension at 72°C for 5 min. PCR products were purified using the QIAquick PCR Purification Kit (Qiagen, Hilden, Germany).
Approximately 100 ng of the purified fragment was cloned into pUPD2 plasmid by GoldenBraid assembly using BsmBI. The resulting constructs were transformed into Escherichia coli TOP10 cells (Thermo Fisher Scientific, USA). Transformants were selected on LB agar supplemented with chloramphenicol (12.5 µg/mL) and X-gal (Thermo Fisher Scientific, USA) for blue - white screening. Plasmid DNA was isolated from white colonies, propagated, and the presence of the correct insert was verified by commercial Sanger sequencing (StarSEQ, Mainz, Germany).
Throughout the study, the plasmid pDB1a1/T7-VNP15 Lime, which shares the same backbone but contains a non-ToBRFV insert, served as the negative control.
| Final concentration in reaction premix (µM) | ||||||
|---|---|---|---|---|---|---|
| Primer name | Primer sequence (5´- 3´) | End-point PCR | LAMP | RPA | Cloning | Designed |
| One-step RTX-PCR | ||||||
| F3_MP1 | TCATAGACTTGTCAAAATCAGAA | 0.1 | Bernabé-Orts. et al., 2022 | |||
| B3_MP1 | GAAGCAAGAGTTGCCTCG | 0.1 | ||||
| FIP_MP1 | GGACAAAGATTCATCTTCATGAACCCGTCTATGTTCACACCTGTT | 0.4 | 0.3 | Bernabé-Orts. et al., 2022 | ||
| BIP_MP1 | AGGTGAATGGAATTTGCCAGATCATTCTCTTATCGACCAAACAG | 0.4 | 0.3 | |||
| LF_MP1 | ACCTTGGAGATCATGACACTCT | 0.2 | Bernabé-Orts. et al., 2022 | |||
| LB_MP1 | TGTCGTGGTGGTGTCAGTG | 0.2 | ||||
| ToBRFV_F3 | TTGGAGTCTTAGATGTTGCG | 0.4 | Rizzo et al., 2021 | |||
| ToBRFV_B3 | GGACACCGTCAACTAGGA | 0.4 | ||||
| GB-PC-ToBRFV-F | GCGCCGTCTCGCTCGAATGCTTTTAAGGAAAAATGTCCATG | 0.3 | this study | |||
| GB-PC-ToBRFV-R | GCGCCGTCTCGCTCAAAGCTCATTGCAAAAGAGATTTGTCA | 0.3 | ||||
| ToBRFV-dF | TTACACAATCGCAACTCCATCGC | 0.5 | this study | |||
| ToBRFV-dR | AGACATATTTAATACGAATCTGAATCGGCG | 0.5 | ||||
Table 1. Primers used in this study. Primers were originally designed by Rizzo et al., 2021, Bernabé-Orts. et al., 2022, or in this study. Cloning primers GB-PC-ToBRFV-F and GB-PC-ToBRFV-R contain target sites for the ++Bsm++BI restriction enzyme (CGTCTC, in bold), four-nucleotide overlaps for cloning into the universal domestication plasmid of the GoldenBraid system pUPD2 (CTCG in the forward, CTCA in the reverse primer, in italics) and a set of four-nucleotide sequences which enables the insertion of the fragment from the universal vector pUPD2 into the infectious potato virus X-clone (AATG in the forward, AAGC in the reverse primer, underlined).
RNA isolation and cDNA synthesis: Total RNA was isolated from approximately 100 mg of plant tissue using RiboEx™ LS (GeneAll® Biotechnology, Seoul, Korea) following the manufacturer’s protocol. RNA pellets were resuspended in 40 µL of DEPC-treated water, and RNA concentration, purity (A260/A280), and spectral profiles were assessed using a NanoDrop spectrophotometer (Thermo Scientific).
For cDNA synthesis, RNA samples were adjusted to a concentration of 500 – 1 000 ng/µL. First-strand cDNA was generated using the RevertAid First Strand cDNA Synthesis Kit with Random Hexamers (Thermo Scientific™, Wilmington, DE, USA). Primer annealing was performed at 65°C for 5 min, followed by reverse transcription at 25°C for 5 min and 42°C for 1 h.
End-point PCR: End-point PCR was performed in 12.5 µL reactions using the Colorless DreamTaq™ Hot-Start PCR Master Mix (Thermo Fisher Scientific, Waltham, MA, USA) with primers ToBRFV_F3 and ToBRFV_B3 (Rizzo et al., 2021; Table 1). Twenty fold diluted cDNA served as the template. Plasmid pDB1α1/PC-ToBRFV/PVX (0.5 ng) was included as the positive control and plasmid pDB1α1/T7-VNP15-Lime (0.5 ng) as the negative control; a no-template control (H2O) was included. Thermal cycling conditions were: 94°C for 5 min; 35 cycles of 94°C for 30 s, 56°C for 30 s, and 72°C for 30 s; followed by a final extension at 72°C for 10 min. PCR products were separated on 2% agarose–TBE gels containing MidoriGreen (NIPPON Genetics EUROPE, Düren, Germany) and visualised using a ChemiDoc™ MP Imaging System (Bio-Rad, Hercules, CA, USA).
One-enzyme RTX-PCR: One-enzyme RTX-PCR, exploiting the combined reverse transcriptase and DNA polymerase activities of RTX, was performed directly from crude plant extracts following Hoffmeisterová et al., 2022. Reactions contained primers ToBRFV_F3 and ToBRFV_B3 (0.4 µM each, Table 1; Rizzo et al., 2021). Plasmid pDB1α1/PC-ToBRFV/PVX (0.5 ng) served as the positive control and pDB1α1/T7-VNP15-Lime (0.5 ng) as the negative control; a no-template control (H2O) was included.
Thermal cycling conditions were: 68°C for 10 min (equivalent of reverse transcription), followed by 33 cycles of 98°C for 10 s, 60°C for 20 s, and 72°C for 20 s, with a final extension at 72°C for 5 min. Amplicons were resolved on 2% agarose–TBE gels and visualised as described above.
Immuno-capture reverse transcription PCR (IC-RTPCR): Immuno-capture (IC) of ToBRFV was performed according to Mulholland (2009) using polyclonal anti-ToBRFV IgG (1:100 in 0.1 M Na2CO3; DSMZ, Braunschweig-Süd, Germany; cat. AS-1236). Positive and negative capture controls consisted of isolated ToBRFV (1 μg/μL) and water, respectively. Reverse transcription was performed directly in the tubes using the RevertAid First Strand cDNA Synthesis Kit (Thermo Scientific™) with Random Hexamers, following the manufacturer’s instructions. No external template was added; water was used to maintain reaction volume. The resulting cDNA was diluted 10-fold and used as template for end-point PCR. Amplicons were separated by agarose gel electrophoresis and visualised as described above.
Loop-mediated isothermal amplification (LAMP) followed by electrophoresis or colorimetric detection using Neutral Red: LAMP assays were performed following Wang et al. (2021) using the Bst 3.0 DNA Polymerase Kit (New England Biolabs, Ipswich, MA, USA). Each 20 μL reaction contained: 2 mM Tris-HCl (pH 8.8), 10 mM (NH4)2SO4, 50 mM KCl, 8 mM MgSO4, 0.8 M betaine, 0.1% Tween 20, primers F3_MP1 and B3_MP1 (0.1 μM each), primers FIP_MP1 and BIP_MP1 (0.4 μM each), primers LF_MP1 and LB_MP1 (0.2 μM each), 20 μg Neutral Red, 8 U Bst 3.0 polymerase, and 1 μL of 20× diluted cDNA. Positive and negative controls consisted of 0.5 ng of plasmid pDB1α1/PC-ToBRFV/PVX and pDB1α1/T7-VNP15-Lime, respectively; a no-template control was included.
Reactions were incubated at 60°C for 40 min followed by enzyme inactivation at 80°C for 5 min. Visual colour assessment distinguished negative (yellow) from positive (orange/pink) reactions. Amplicons were confirmed by electrophoresis on 2% agarose–TBE gels and visualized as described above.
Recombinase polymerase amplification (RPA): RPA was carried out using the Lyo-Ready RPA Kit (Invitrogen™, Thermo Fisher Scientific, Waltham, MA, USA) according to the manufacturer’s instructions using 0.3 μM of each primer (FIP_MP1 and BIP_MP1) and 0.5 μL 20× diluted cDNA as template per 20 μL reaction. Positive and negative controls consisted of 0.5 ng of plasmid pDB1α1/PC-ToBRFV/PVX and pDB1α1/T7-VNP15-Lime, respectively, and a no-template control was included. Reactions were incubated at 40°C for 20 min in a dry block heater and subsequently cooled on ice. Products were analysed on 2% agarose–TBE gels and visualized as described above.
SDS-PAGE and Coomassie Brilliant Blue R-250 staining: Isolated ToBRFV preparations were mixed with standard SDS-PAGE sample buffer containing 6 M urea and incubated at 65°C for 20 min. Samples were resolved on 15% polyacrylamide gels following the method of Laemmli (1970). Gels were rinsed shortly three times with water, stained overnight with Coomassie Brilliant Blue R-250, and destained according to Brunelle and Green (2014).
SDS-PAGE and Western blot: Following SDS-PAGE, proteins were electroblotted onto a nitrocellulose membrane (Amersham™ Protran™ Premium, 0.45 μm pore size; Cytiva, Wilmington, DE, USA) at a constant current density of 0.8 mA/cm² for 1.5 h, following the procedure of Towbin et al. (1979). After the transfer, the membrane was rinsed three-times for 5 min in distilled water, and the target proteins were subsequently detected immunochemically as described below.
Tissue blot immunoassay (TBIA): TBIA was carried out essentially as described by Lin et al. (1990) and Hoffmeisterová et al. (2025). Systemic leaves from ToBRFV-infected tomato plants were cut with a razor blade and gently pressed onto a nitrocellulose membrane (Amersham™ Protran™ Premium, 0.45 μm; Cytiva) for approximately 2 s. In all cases, leaves from healthy tomato plants served as negative controls, whereas 1 μL of isolated ToBRFV (1 μg/μL) was pipetted (dotted) onto the membrane as a positive control. Membranes were airdried for at least 20 min prior to further processing.
The main experiment comprised 14 inoculated and 14 uninoculated control tomato plants. Tissue printing was performed at 7th, 14th, 21st and 28th dpi. Two experimental layouts were used: a) combined layout – all plants (including controls) were printed onto a single membrane at each sampling point; b) individual layout – each plant, together with one of negative controls, was printed on a separate membrane at each time point to visualize the progression of infection.
For samples collected from the commercial greenhouse in Chornice, seven-month-old tomato leaves were tissueprinted on site by greenhouse personnel, and membranes were subsequently processed at the Czech Agrifood Research Center (CARC). Leaves from five-month-old tomato plants grown in private gardens or small greenhouses were transported to the CARC under cooled conditions within 24 h, tissue-printed and processed.
Seed transmission was evaluated by printing stems of four-week-old seedlings grown from seeds harvested from ToBRFV-infected plants.
Dot blot (DB): Dot blot analysis followed by immunochemical detection of ToBRFV was performed on isolated virus preparations and on extracts from tomato seeds collected from infected plants, following Hoffmeisterová et al. (2025). Subsequent immunodetection of ToBRFV was carried out as described below. One microlitre of purified ToBRFV (1 μg/μL) served as a positive control, whereas healthy plants were tissueprinted as negative controls.
Immunochemical detection using polyclonal anti-ToBRFV antibodies: Following Western blotting, tissue printing or dot blotting, membranes were immunodetected according to Hoffmeisterová et al. (2025). Primary polyclonal anti-ToBRFV antibodies (1:1 000 in PBS containing 0.05% Tween 20; DSMZ, Leibniz Institute, Braunschweig-Süd, Germany) and secondary goat antirabbitIgG antibodies conjugated with alkaline phosphatase (1:20 000 in PBS containing 0.05% Tween 20; Sigma-Aldrich, Merck, Darmstadt, Germany) were used. Final signal visualization was carried out using SigmaFAST™ BCIP®/NBT substrate (Sigma-Aldrich, Merck, Darmstadt, Germany).
Tomato single-seed ToBRFV detection system: A single tomato seed was thoroughly ground in liquid nitrogen, followed by the addition of 0.5 mL of CTAB extraction buffer (2% CTAB, 2% PVP-40, 100 mM Tris-HCl pH 8.0, 1.4 M NaCl, 20 mM EDTA, and 0.2% β-mercaptoethanol). The mixture was incubated at 65°C for 45 - 60 min and centrifuged at 13 000 × g for 10 min. A total of 450 μL of the supernatant was transferred to a new 1.5 mL microcentrifuge tube and mixed with an equal volume of chloroform/isoamyl alcohol (24:1) by vortexing for 30 s. After centrifugation at 13 000 × g for 10 min, 300 μL of the aqueous phase was carefully transferred to a fresh 1.5 mL tube and mixed with 250 μL of isopropanol. The tube was gently inverted to mix and incubated at room temperature for 10 min, followed by centrifugation at 13 000 × g for 15 min. The supernatant was removed, and the pellet was washed with 500 μL of 75% ethanol and centrifuged at 13 000 × g for 10 min. After removing the supernatant, the pellet was air-dried at room temperature for 5 min. Finally, the pellet was resuspended in 40 μL of RNasefree water and incubated on ice for 30 min with occasional pipetting to facilitate dissolution. The extracted nucleic acids were stored at –80°C until use. cDNA synthesis was performed using the M-MLV Reverse Transcriptase Kit (Promega, Madison, WI, USA) according to the manufacturer’s instructions. The synthesized cDNA was stored at –20°C until use. PCR was conducted using the Taq-Plus DNA Polymerase Kit (Sangon Biotech, Shanghai, China) according to the manufacturer’s instructions using 0.5 μM each of primers ToBRFV-dF and ToBRFV-dR and 1 μL of undiluted cDNA as template per 20 μL reaction. The PCR conditions consisted of an initial denaturation at 94°C for 3 min; 35 cycles of 94°C for 30 s, 56°C for 30 s, and 72°C for 1 min; followed by a final extension at 72°C for 10 min. Products were separated on agarose gel with an addition of ethidium bromide and visualized using GenoSens 1850 gel imaging system (Shanghai Qinxiang, Shanghai, China).
Results
ToBRFV propagation and isolation: ToBRFV was readily and rapidly propagated in both N. benthamiana and tomato, with typical symptoms including mosaic, leaf deformation, rugosity, and chlorosis along veins appearing around 14 dpi in N. benthamiana and around 21 dpi in tomato (Fig. 1). The isolation protocol for ToBRFV from N. benthamiana leaves yielded high amounts of the virus (approximately 40 mg per 100 g of infected leaves) and ensured high purity and reactivity of the isolated virus (Fig. 3A1-A3).
Molecular methods: For the evaluation of molecular detection methods for ToBRFV, eight tomato plants were inoculated, while two plants were left uninoculated as healthy controls. All inoculated plants developed characteristic symptoms at 21 dpi, whereas two healthy plants showed no symptoms (Fig. 1). Samples from upper/systemic leaves were collected from all plants at 21st dpi and stored at –80°C until analysis.
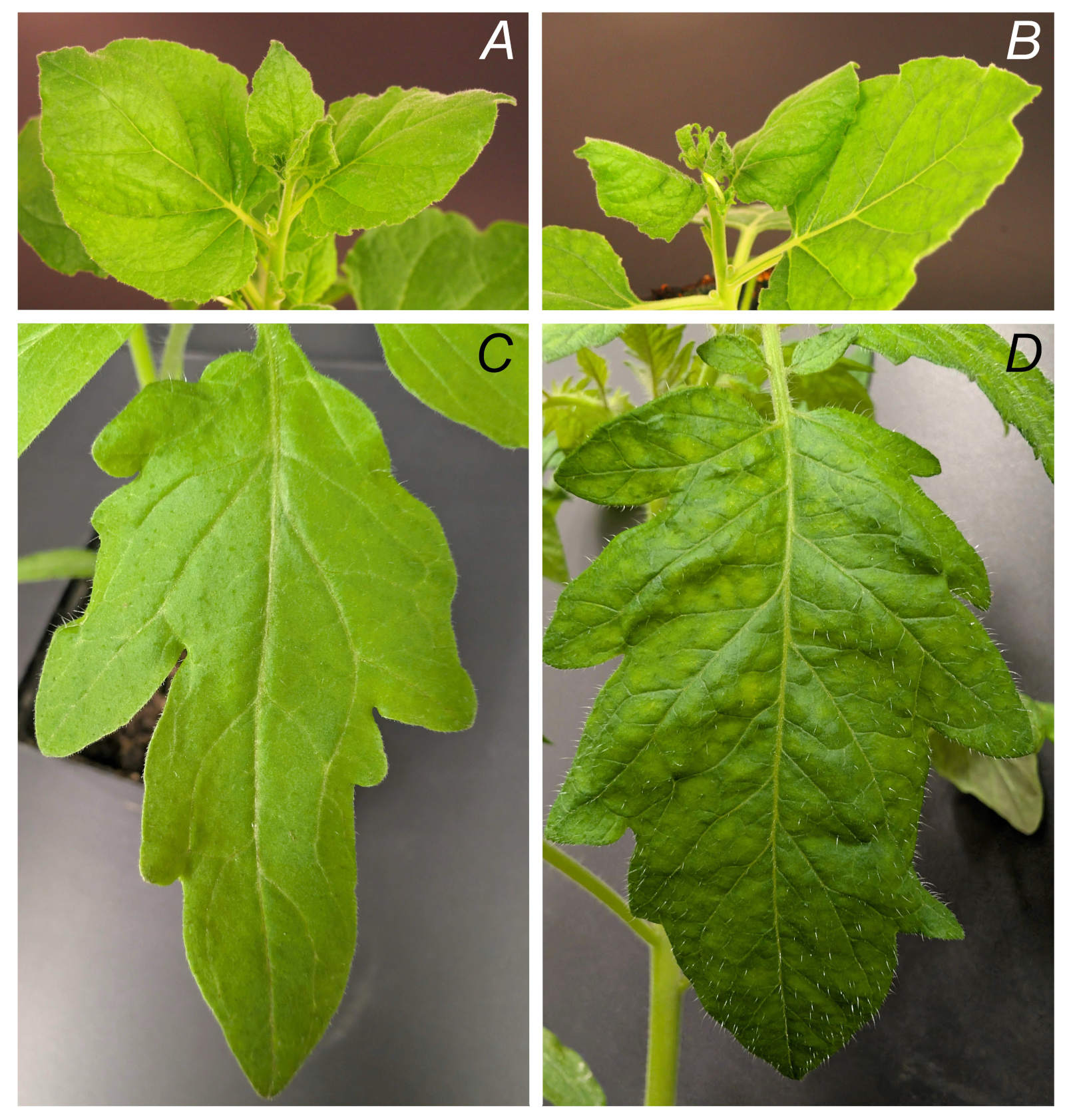

Fig. 1. Symptoms of ToBRFV. A - healthy N. benthamiana, B - infected N. benthamiana (14 dpi), C - healthy tomato cv. Gallant; D - infected tomato cv. Gallant (21 dpi).
Using RT-PCR, all inoculated plants and also the positive control were detected as positive as a band of the correct size of 279 bp was detected on gel. In uninfected plants as well as in negative controls, namely healthy tomato plants and plasmid containing no target sequence, no signal/band was detected (Fig. 2A).
When performing one-enzyme RTX-PCR and the same templates and primers as conventional RT-PCR, analogous results to RT-PCR were obtained which means all inoculated plants and the positive control were tested as positive showing the band of 279 bp, the negative controls and uninfected plants as negative (Fig. 2B) showing no band.
Immunocapture RT-PCR (IC-RT-PCR) using for the final step the same primers as RT-PCR and one-enzyme RTX-PCR gives also the analogous result which means that all infected tomato plants as well as the positive control showed the band of correct size of 279 bp, while negative tomatoes and the negative controls showed no band on electrophoresis (Fig. 2C).
When using RPA, the product of expected size was detected in all samples of inoculated plants and the positive control, no bands were detected in healthy plants and negative controls (Fig. 2D).
A loop-mediated isothermal amplification (LAMP) reaction using six primer pairs and the neutral red dye resulted in a colour change from yellow to pink/orange in all samples of inoculated plants, as well as in the positive control. Meanwhile, the colour of the reaction mixture remained unchanged (yellow) in the healthy plants and negative controls (Fig. 2F). After the reaction mixtures were loaded onto a gel, a typical LAMP product pattern of bands of different sizes was observed in all inoculated plant samples and the positive control. No bands were detected in the samples of healthy plants and the negative controls (Fig. 2E).
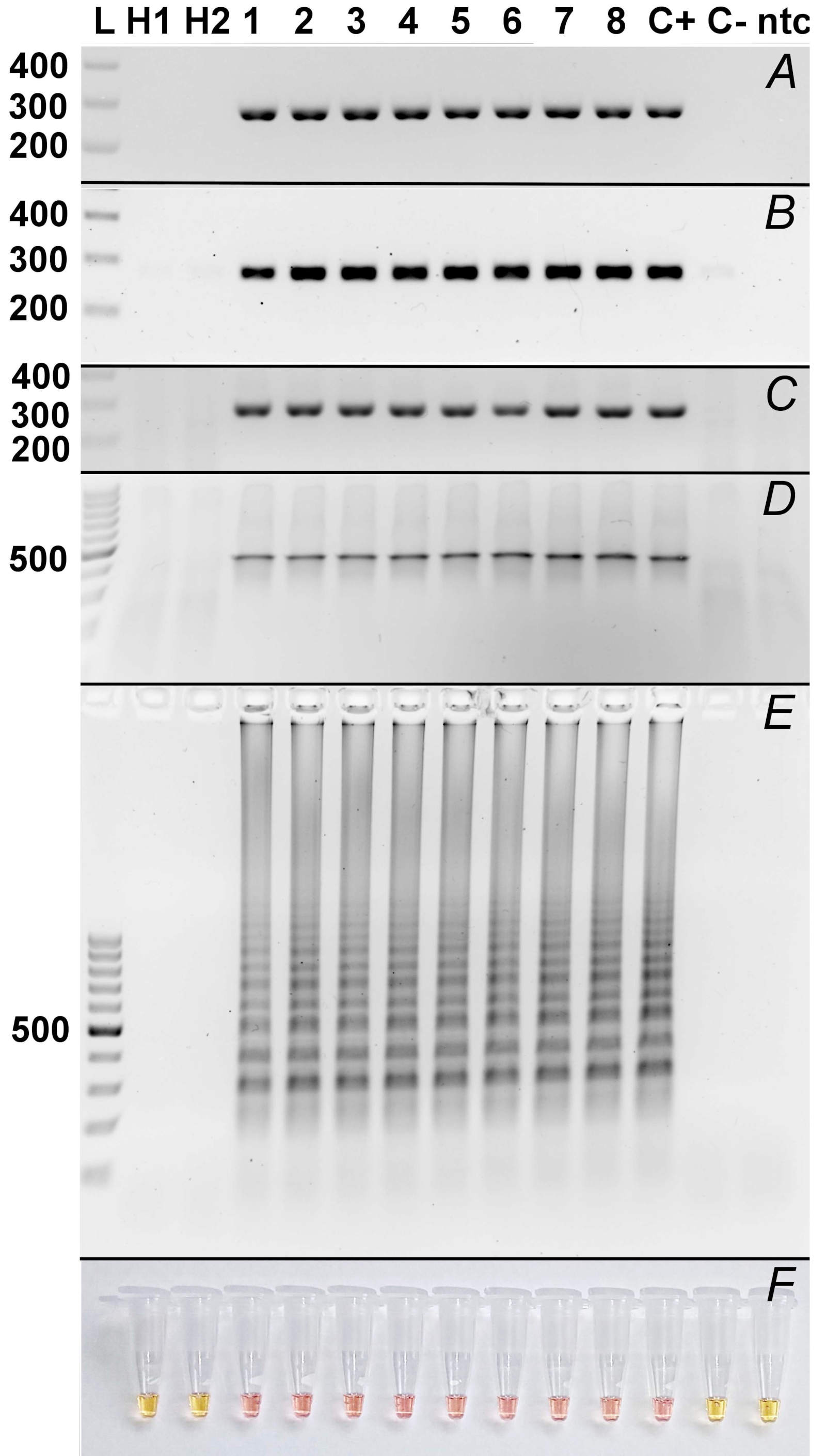
Fig. 2. Comparison of molecular methods used for detection of ToBRFV. A - RT-PCR; B - One-enzyme RTX-PCR; C - IC-RT-PCR; D - Lyo-Ready RPA; E - LAMP; F - LAMP – colorimetric detection using Neutral Red. L: Thermo ScientificTM GeneRuler 100 bp DNA Ladder (Thermo Fisher Scientific, Waltham, MA, USA); H1, H2: healthy plant no. 1, 2; 1 - 8: infected plants no. 1 - 8; C+: positive control (ToBRFV-pDB1α1/PC-ToBRFV/PVX plasmid); C-: negative control (pDB1a1/T7-VNP15-Lime plasmid); ntc: no-template control.
Serological methods: For immunodetection, ToBRFV-specific antibodies were evaluated using Western blot, DB, and TBIA assays. The antibodies exhibited strong and specific reactivity, as shown in Fig. 3A2 and A3.
Fourteen infected and fourteen healthy tomato plants were tested in two parallel tissue-blot immunoassay (TBIA) layouts. In the first layout, all plants, including controls, were tissue-printed onto a single membrane at 7th, 14th, 21st, and 28th dpi. In the second layout, each plant and a single control plant were tissue-printed separately onto individual membranes on each processing day to monitor infection progression over time. As demonstrated in Fig. 3B, all infected plants at 14th dpi as well as positive control showed positive signals, whereas all healthy plants remained negative. Evidence of infection progression over time was estimated according to the increasing signal on the immunoprocessed nitrocellulose membrane in inoculated plant while no positive signal was detected in healthy plant till the 28th dpi (Fig. 3C).
Monitoring of ToBRFV in commercial greenhouses and private gardens and field plots: All 73 samples collected from the Chornice commercial greenhouse between 2023 and 2025 were tested as negative by TBIA (Fig. 3D, 60 samples shown) and one-enzyme RTX-PCR (data not shown). Similarly, all 131 samples from private gardens and field plots across various locations in the Czech Republic during the same period were negative (data not shown).
Vertical transmission of ToBRFV in tomato: Dot-blot analysis with immunochemical detection was performed on tomato seeds. Twenty-one out of 43 seed samples tested positive by DB and subsequent immunochemical detection (Fig. 3E). However, none of the plants grown from seeds harvested from ToBRFV-infected plants tested positive using either TBIA or one-enzyme RTX-PCR (data not shown).
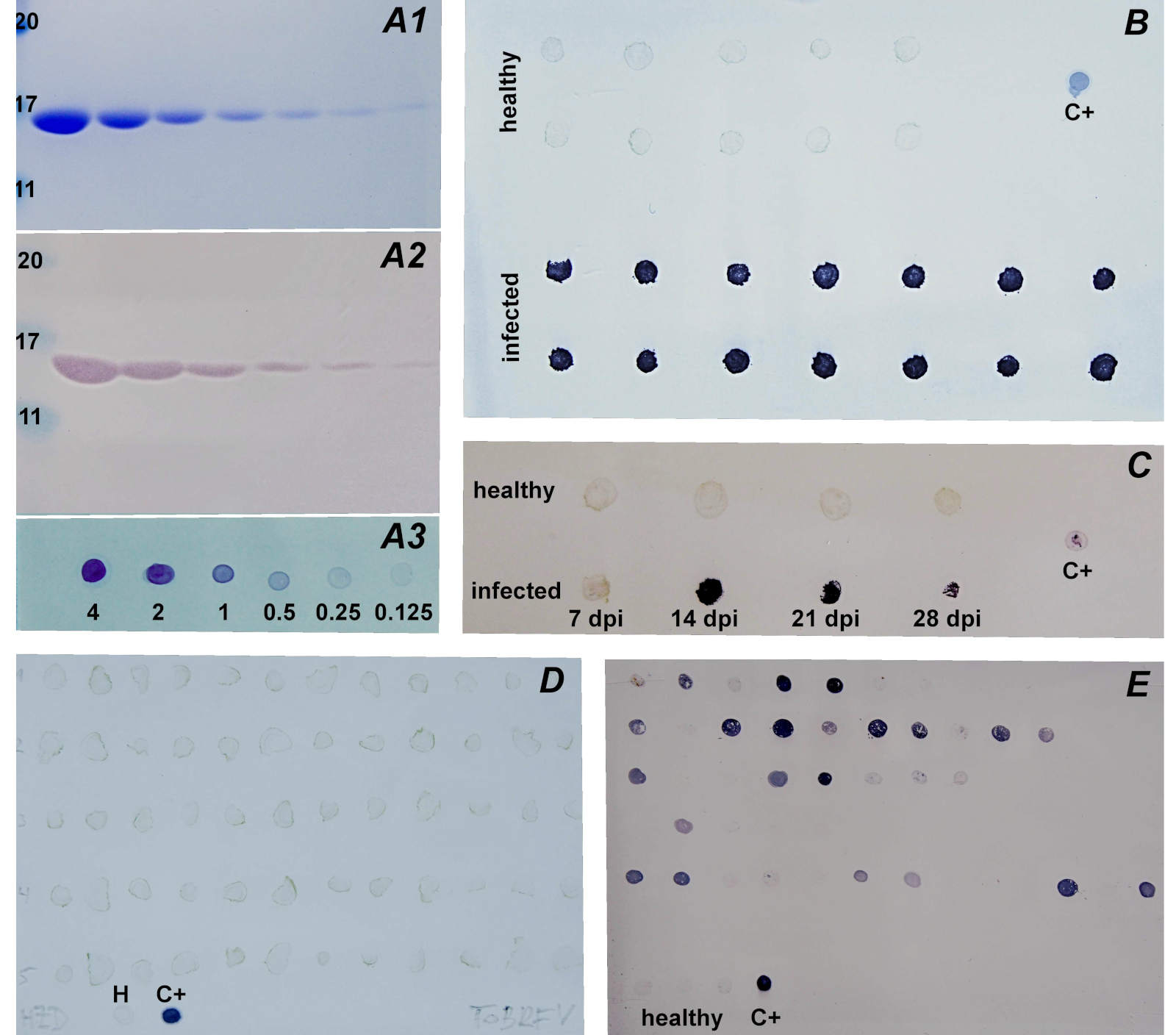
Fig. 3. Protein and immunochemical detection of ToBRFV using anti-ToBRFV antibodies. A - detection of isolated and diluted ToBRFV, numbers below mean amounts of ToBRFV (μg), numbers on the left mean protein size (kDa). A1 - SDS-PAGE/Coomassie Brilliant Blue R-250 detection; A2 - SDS-PAGE/Western blot; A3 - Dot-blot. B - TBIA of healthy and infected tomato plants at 14th dpi. C - TBIA of one healthy and one infected plant at different dpi. D - TBIA of 60 of 73 in total tomato plants grown commercially in Chornice greenhouses. E - Dot-blot of samples of seeds harvested from ToBRFV-infected plants, each individual row represents seeds from one individual fruits. H: healthy plant. C+: isolated ToBRFV (1 μg).
Single-seed ToBRFV detection system in tomato: A ToBRFV detection method for single tomato seeds was developed. The tomato seeds collected from ToBRFV infected and healthy plants were individually tested for ToBRFV using RT-PCR. All seeds harvested from infected plants showed the expected amplification of a 472 bp product, indicating the presence of ToBRFV. In contrast, no target band was amplified in samples of seed obtained from healthy plant (Fig. 4).
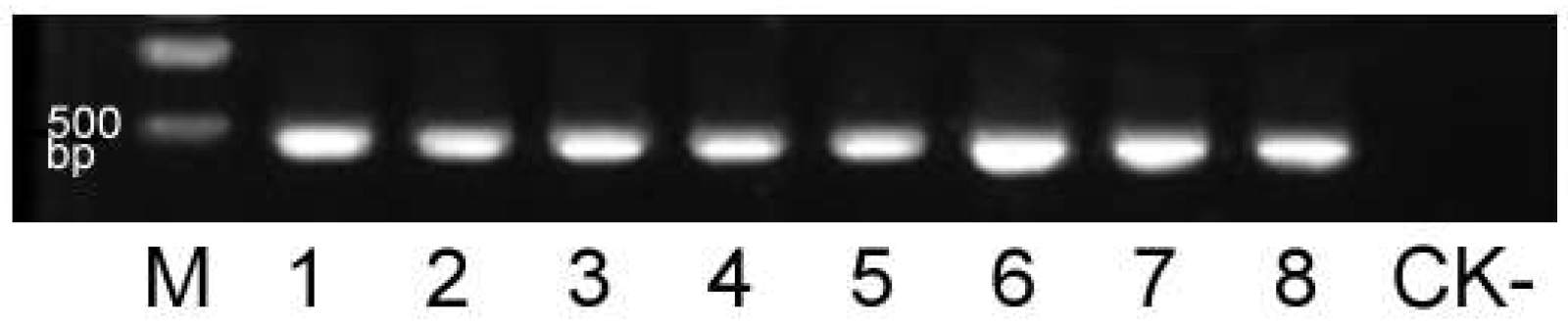
Fig. 4. RT-PCR detection of ToBRFV in single tomato seeds. M: DNA marker (Trans2K Plus DNA Marker, Beijing Quanshijin Biotechnology Co., Ltd., Beijing, China); lanes 1 - 8: individual seeds harvested from ToBRFV-infected tomato plants; CK-: seed from a healthy tomato plant.
Discussion
Tomato brown rugose fruit virus (ToBRFV) is a tobamovirus, according to the EPPO reports, first detected in Israel in 2014, and soon after in Jordan, causing severe fruit rugosity and yield loss in tomatoes and peppers. Because it is transmissible easily by plant contact and contaminated tools (Caruso et al., 2022; Salem et al., 2022; Esmaeilzadeh et al., 2023), the virus rapidly spread to Europe, America, and Asia within just a few years (EPPO, 2026). Its ability to break common tobamovirus resistance genes such as Tm-2² alarmed growers and regulators worldwide (Luria et al., 2018; Maayan et al., 2018). Therefore, in 2019, the European Union and EPPO listed ToBRFV as a quarantine pest, followed by the United States and other regions worldwide. Today, ToBRFV is a quarantine pathogen in Morocco and China and A1 or A2 list-pathogen in several countries all around the world (EPPO, 2020a). Since then, strict seed testing, hygiene protocols, and movement controls have been implemented to limit further outbreaks and protect global Solanaceous crop production. At the same time, there was a strong need for quick, reliable, easily performable, highly specific, and sensitive detection methods. In the present study, we compare several molecular and serological methods for the detection of ToBRFV. Next, we describe the preparation of safe positive controls for molecular methods, which is crucial for safe handling. Finally, we developed a tomato single-seed assay to test individual seeds directly from seed stocks.
For the initial propagation of ToBRFV, N. benthamiana plants were used. As expected, all plants became severely infected after mechanical inoculation because ToBRFV belongs to the Tobamovirus family, whose members infect host plants very easily (Dawson, 1992). Due to the relatedness of ToBRFV and TMV, the protocol for isolating TMV was successfully used to isolate ToBRFV. After mechanical inoculation, TMV probably reaches the highest virus titer in plant tissues among plant viruses (Lomonossoff and Wege, 2018), with a yield of 40 - 141 mg per 100 g of N. benthamiana leaves (Monroy-Borrego and Steinmetz, 2022). In this study, the ToBRFV yield from infected N. benthamiana leaves was estimated at 40 mg per 100 g, fitting into the TMV yield range. The purity of the isolated ToBRFV (Fig. 3A1) confirms the protocol’s suitability for virus isolation.
The implementation of safe positive controls, such as synthetic RNA standards and recombinant plasmids of quarantine organisms, including viruses, enhances biosafety and lowers environmental risks, especially in laboratories lacking high-containment facilities. Therefore, we designed several plasmids containing highly conserved regions of ToBRFV replicase or movement protein (MP). All plasmids/positive controls reacted successfully with the appropriate primers (data not shown) when used in molecular methods. To obtain a safe positive control for serological methods as well, we attempted to express constructs containing ToBRFV MP sequences using PVX- or apple latent spherical virus (Cheravirus mali, ALSV)-based viral vectors in N. benthamiana via Agrobacterium-mediated transient expression. Unfortunately, we repeatedly detected only empty viral vectors (wild-type PVX or ALSV), most likely referring to the loss of the insert during expression in plants, which occurs quite often (Avesani et al., 2007; Chung et al., 2007). However, even if the insert is stable, expression of the recombinant protein can be unsuccessful (Igarashi et al., 2009). For our experiments described throughout this publication, we finally decided to use one of the positive controls applicable in molecular methods – ToBRFV-pDB1α1/PC-ToBRFV/PVX, a construct inserted into a PVX-based viral vector, with which the Laboratory of Virology has long-term experience (Čeřovská et al., 2008; Cerovska et al., 2009; 2012; Plchova et al., 2011; Plchová et al., 2022).
For the comparison of molecular and immunological methods throughout this paper, two healthy and eight ToBRFV-mechanically inoculated tomato cv. Gallant plants were used. Tomato cv. Gallant was earlier genotyped for tobamovirus resistance loci and was found to carry the classic Tm‑2² allele, which is widely deployed for resistance to tobamoviruses such as TMV/ToMV. However, ToBRFV has been reported to overcome Tm‑2²-mediated resistance, and in our recent marker assays, tomato cv. Gallant did not show the resistant-allele patterns for Tm‑1/Tm‑2 associated with ToBRFV resistance (Ibrahim et al., 2026). The experiment was repeated three times, and in all repetitions, all inoculated plants became infected and tested positive (Fig. 3). This result was expected because of the chosen pathogen–host pair, i.e., tobamovirus–highly susceptible tomato plants (Ershova et al., 2023).
Among molecular methods, we tested RT-PCR, one-enzyme RTX-PCR, IC-RT-PCR, RPA, and LAMP followed by both electrophoretic and colorimetric detection. All molecular methods are highly specific and more sensitive than serological methods; however, the required equipment is usually expensive, and only well-trained staff can handle the samples properly (Olmos et al., 2007; Jeong et al., 2014; Cassedy et al., 2021). In comparison, serological methods such as Western blot, dot blot, and TBIA are less specific and sensitive; nevertheless, in some cases, untrained staff can handle the samples, even directly in field conditions. Furthermore, some serological methods, such as dot blot or TBIA, can process tens or hundreds of samples easily and quickly, usually without any need for expensive instruments (Torrance, 1997; Al-Moslih, 2012).
As mentioned previously, for the molecular detection of the virus, we used classical RT-PCR, one-enzyme RTX-PCR, RPA, and LAMP followed by electrophoretic and colorimetric detection. Conventional RT-PCR represents a routinely proven method. Prior to the PCR itself, RNA isolation and subsequent reverse transcription (RT) are needed. RNA isolation is time-consuming and, especially when performed using column kits, quite expensive. One-enzyme RTX-PCR using the RTX reverse transcriptase/DNA polymerase (Hoffmeisterová et al., 2022) offers several advantages in comparison with RT-PCR. First, convenience, since RT and PCR run in the same tube as a one-step reaction; second, speed, because the RTX enzyme performs RT efficiently, typically within 5 - 10 min; third, complete omission of RNA isolation, since crude plant extract supernatant can be used directly as template; fourth, because of the use of a home-made RTX enzyme, the performance is even more cost-effective. Immunocapture RT-PCR also skips the RNA isolation step, since virions are captured using specific antibodies, which, as noted below, crossreact with some other tobamoviruses. However, at the PCR step, specificity is ensured using specific primers. For the RPA, we used commercial kits that used either RNA or cDNA as templates. The kit that uses RNA as a template ensures a one-tube reaction, but it is naturally more expensive than the kit that uses cDNA as a template. In the latter case, however, the RT step is required prior to RPA. The unquestionable advantage is the isothermal amplification, which can be run in a low-cost dry block heater instead of a thermocycler. The LAMP method, combined with colorimetric detection, uses three pairs of primers whose concentrations need to be optimised. Its advantage arises from the visual colour change, allowing samples to be sorted easily according to their colours without electrophoresis; however, convenient gel electrophoresis can be used to analyse the reaction products if needed.
Before immunodetection of ToBRFV, the specificity of the anti-ToBRFV antibodies among several viruses belonging to the genus Tobamovirus – namely TMV, tomato mosaic virus (Tobamovirus tomatotessellati, ToMV), tobacco mild green mosaic virus (Tobamovirus mititessellati, TMGMV), and pepper mild mottle virus (Tobamovirus capsici, PMMV), all available in the collections of CARC and/or IEB – were tested. Crossreaction of the ToBRFV antibodies with ToMV and TMV was observed (data not shown), meaning the antibodies do not fully ensure specificity. Therefore, in cases of immunologically positive samples, we recommend verifying the presence of ToBRFV using one of the molecular methods.
For immunodetection, SDS-PAGE/Western blot represents a conventional method for testing samples.This method provides not only information about sample positivity but also indicates potential cross-reactivity with plant proteins. SDS-PAGE/Western blot is, however, limited by the number of samples that can be processed in the same day – usually not more than a few dozen. We used this method for testing isolated ToBRFV to assess antibody reactivity, which was estimated down to tens of nanograms of the target proteins (64 ng). The anti-ToBRFV antibodies did not cross-react with any N. benthamiana or tomato proteins (data not shown). Therefore, dot blot (DB) could also be used. DB is suitable for testing hundreds of samples in a batch. Prior to testing, plant samples are crushed in a suitable buffer, centrifuged, and the supernatant is dotted onto nitrocellulose membranes, which are subsequently immunoprocessed. Because the used antibodies do not cross-react with plant proteins, DB is appropriate; however, because of cross-reactivity with several tobamoviruses, molecular confirmation is recommended. We successfully used DB, particularly for seed testing, since TBIA is more difficult to perform on seeds.
Tissue blot immunoassay (TBIA) was used as a quick and easy method in which sample transfer onto membranes is performed simply by printing cut stems/petioles – something that can be done directly in the field or greenhouse. In our experiment, 14 infected and 14 healthy tomato plants were printed, either to observe overall infection or to follow infection progression over time. After printing, ToBRFV was immunodetected. As shown in Fig. 3B, all infected and none of the healthy plants showed a positive signal. The progression of infection and the related increase in virus titer in the early stages of infection (7 dpi and 14 dpi) was confirmed (Fig. 3C). Over time, the virus titer decreased, however, at 28 dpi, the amount of virus detected was still higher than in the positive control (1 μg of isolated ToBRFV). The first symptoms occurred about one week after ToBRFV was immunochemically detected (14 dpi). Although TBIA is not a quantitative method, the rapid increase in virus titer at 14 dpi followed by a slow decrease over time appears to follow the general pattern of virus titer during infection, as observed in other virus–host systems (Hewings et al., 1990; Lapidot et al., 1997; Bachand and Castello, 1998). For screening tomatoes grown in the commercial greenhouse in HZDJ in Chornice, greenhouse staff tissue-printed random leaves from approximately sevenmonth-old plants onto membranes, which were then delivered to CARC for immunoprocessing. Leaves from approximately five-month-old random tomato plants grown in private gardens and greenhouses were collected, kept cool, and delivered to CARC within one day, where they were tissue-printed and immunoprocessed. No positive samples were detected in Chornice greenhouses (Fig. 3D) or in the privately grown tomatoes (data not shown), indicating current negligible ToBRFV distribution in the Czech Republic, even under field conditions. In the past, according to the European and Mediterranean Plant Protection Organization (EPPO) report, the first occurrence of ToBRFV in Czech Republic was in August 2020 (EPPO, 2020b). The virus was detected in a greenhouse of a breeding station of a seed company during pre-export testing of a Capsicum annuum (pepper) seed crop. Following that initial detection, later official findings did include Solanum lycopersicum (tomato) fruit production in the Ústecký region (November 2020) and in the Jihomoravský region (January 2021), and a pepper seed company in the Moravskoslezský region (October 2021).
To test the vertical transfer of ToBRFV in tomato plants, we performed DB analysis of seeds harvested from infected plants, followed by immunochemical detection of ToBRFV; 21 samples out of 43 were considered positive (Fig. 3E). However, no plant grown from 47 seeds harvested from ToBRFV-infected plants tested positive using TBIA or one-enzyme RTX-PCR. The published ToBRFV seed transmission rate is 2.8% in cotyledons and 1.8% in the third true leaf (Dall et al., 2023). In our case, we obtained only 90 seeds in total from ToBRFVinfected plants, approximately half of which were tested by the destructive DB method, while the rest were sown. The combination of low ToBRFV transmission rate and low number of tested plants can explain why all tested seedlings were ToBRFV-negative.
Several papers have reported detecting ToBRFV or other tobamoviruses in tomato seeds, but all of them tested for the presence of the virus in whole batches of seeds, not in individual seeds (Davino et al., 2020; Rizzo et al., 2021; Vargas-Hernández et al., 2022; Dall et al., 2023). To test individual tomato seeds in seed stocks, we developed a tomato single-seed assay that enables accurate detection of ToBRFV contamination in individual seeds.
In conclusion, this study demonstrates that a combination of molecular and serological diagnostic approaches offers the most robust framework for detecting tomato brown rugose fruit virus (ToBRFV) across diverse experimental and practical settings. Our results underscore the importance of integrating complementary molecular and serological diagnostics to achieve reliable, sensitive, and practical detection of ToBRFV. Such an integrated strategy is essential for effective surveillance, containment, and phytosanitary management of this economically significant quarantine virus in global tomato and pepper production systems.
Download citation
References
- Al-Moslih, M. (2012) Serological Diagnosis of Certain Human, Animal and Plant Diseases. London: IntechOpen, pp. 172.
 Go to original source...
Go to original source... - Asselin, A. & Zaitlin, M. (1978) Characterization of a second protein associated with virions of tobacco mosaic virus. Virology, 91, 173-181.
 Go to original source...
Go to original source... - Avesani, L., Marconi, G., Morandini, F. et al. (2007) Stability of Potato virus X expression vectors is related to insert size: implications for replication models and risk assessment. Transgenic Research, 16, 587-597.
 Go to original source...
Go to original source... - Bachand, G.D. & Castello, J.D. (1998) Seasonal pattern of tomato mosaic tobamovirus infection and concentration in red spruce seedlings. Applied and Environmental Microbiology, 64, 1436-1441.
 Go to original source...
Go to original source... - Bernabé-Orts, J.M., Hernando, Y. & Aranda, M.A. (2022) Toward a CRISPR-based point-of-care test for tomato brown rugose fruit virus detection. PhytoFrontiers, 2, 92-100.
 Go to original source...
Go to original source... - Brunelle, J.L. & Green, R. (2014) Coomassie blue staining. Methods in Enzymology, 541, 161-167.
 Go to original source...
Go to original source... - Caruso, A.G., Bertacca, S., Parrella, G., Rizzo, R., Davino, S. & Panno, S. (2022) Tomato brown rugose fruit virus: A pathogen that is changing the tomato production worldwide. Annals of Applied Biology, 181, 258-274.
 Go to original source...
Go to original source... - Cassedy, A., Parle-McDermott, A. & O'Kennedy, R. (2021) Virus detection: a review of the current and emerging molecular and immunological methods. Frontiers in Molecular Biosciences, 8, 637559.
 Go to original source...
Go to original source... - Cerovska, N., Hoffmeisterova, H., Moravec, T. et al. (2012) Transient expression of Human papillomavirus type 16 L2 epitope fused to N- and C-terminus of coat protein of Potato virus X in plants. Journal of Biosciences, 37, 125-133.
 Go to original source...
Go to original source... - Cerovska, N., Plchova, H., Moravec, T. et al. (2009) Transient expression of E7 and L2 derived peptides from human papillomavirus type 16 using Potato virus X-based vector. FEBS Journal, 276, 275.
- Čeřovská, N., Hoffmeisterová, H., Pečenková, T. et al. (2008) Transient expression of HPV16 E7 peptide (aa 44-60) and HPV16 L2 peptide (aa 108-120) on chimeric potyvirus-like particles using Potato virus X-based vector. Protein Expression and Purification, 58, 154-161.
 Go to original source...
Go to original source... - Chung, B.-N., Canto, T. & Palukaitis, P. (2007) Stability of recombinant plant viruses containing genes of unrelated plant viruses. Journal of General Virology, 88, 1347-1355.
 Go to original source...
Go to original source... - Dall, D.J., Lovelock, D.A., Penrose, L.D.J. & Constable, F.E. (2023) Prevalences of Tobamovirus contamination in seed lots of tomato and capsicum. Viruses, 15, 883.
 Go to original source...
Go to original source... - Davino, S., Caruso, A.G., Bertacca, S., Barone, S. & Panno, S. (2020) Tomato brown rugose fruit virus: seed transmission rate and efficacy of different seed disinfection treatments. Plants, 9, 1615.
 Go to original source...
Go to original source... - Dawson, W.O. (1992) Tobamovirus-plant interactions. Virology, 186, 359-367.
 Go to original source...
Go to original source... - Dombrovsky, A. & Smith, E. (2017) Seed transmission of tobamoviruses: aspects of global disease distribution. In: Jimenez-Lopez, J.C. (Ed.) Advances in Seed Biology. London: IntechOpen, pp. 231-256.
 Go to original source...
Go to original source... - Dusek, J., Plchova, H., Cerovska, N. et al. (2020) Extended set of GoldenBraid compatible vectors for fast assembly of multigenic constructs and their use to create geminiviral expression vectors. Frontiers in Plant Science, 11, 522059.
 Go to original source...
Go to original source... - EPPO (2020a) EPPO Reporting Service no. 10 - 2020. New additions to the EPPO A1 and A2 Lists. Available at: https://gd.eppo.int/reporting/article-6887.
- EPPO (2020b) EPPO Reporting Service no. 10 - 2020. First report of tomato brown rugose fruit virus in the Czech Republic. Available at: https://gd.eppo.int/reporting/article-6901.
- EPPO (2026) Tobamovirus fructirugosum (TOBRFV). Available at: https://gd.eppo.int/taxon/TOBRFV/reporting.
- Ershova, N., Kamarova, K., Sheshukova, E., Antimonova, A. & Komarova, T. (2023) A novel cellular factor of Nicotiana benthamiana susceptibility to tobamovirus infection. Frontiers in Plant Science, 14, 1224958.
 Go to original source...
Go to original source... - Esmaeilzadeh, F., Santosa, A.I., Çelik, A. & Koolivand, D. (2023) Revealing an Iranian isolate of tomato brown rugose fruit virus: complete genome analysis and mechanical transmission. Microorganisms, 11, 2434.
 Go to original source...
Go to original source... - Hewings, A.D., Damstreegt, V.D., Sindermann, A.E. & Tolin, S.A. (1990) Variation in serologically detectable antigen of soybean dwarf virus in soybean leaflets as a function of time after inoculation and plant age. Plant Disease, 74, 844-848.
 Go to original source...
Go to original source... - Hoffmeisterová, H., Ibrahim, E., Liu, Q. et al. (2025) Efficient serological and molecular methods for the detection of tomato spotted wilt virus. Biologia Plantarum, 69, 77-87.
 Go to original source...
Go to original source... - Hoffmeisterová, H., Kratochvílová, K., Čeřovská, N. et al. (2022) One-enzyme RTX-PCR for the detection of RNA viruses from multiple virus genera and crop plants. Viruses, 14, 298.
 Go to original source...
Go to original source... - Ibrahim, E., Hoffmeisterová, H., Bharati, R. & Kundu, J. (2026) Assessment of tomato cultivar responses to tomato brown rugose fruit virus and tomato spotted wilt virus. Crop and Pasture Science, CP25312.
- Igarashi, A., Yamagata, K., Sugai, T. et al. (2009) Apple latent spherical virus vectors for reliable and effective virus-induced gene silencing among a broad range of plants including tobacco, tomato, Arabidopsis thaliana, cucurbits, and legumes. Virology, 386, 407-416.
 Go to original source...
Go to original source... - Islam, W., Naveed, H., Zaynab, M., Huang, Z. & Chen, H.Y.H. (2019) Plant defense against virus diseases; growth hormones in highlights. Plant Signaling and Behavior, 14, 1596719.
 Go to original source...
Go to original source... - Jeong, J.-J., Ju, H.-J. & Noh, J. (2014) A review of detection methods for the plant viruses. Research in Plant Disease, 20, 173-181.
 Go to original source...
Go to original source... - Laemmli, U.K. (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature, 227, 680-685.
 Go to original source...
Go to original source... - Lapidot, M., Paran, I., Ben-Joseph, R. et al. (1997) Tolerance to cucumber mosaic virus in pepper: development of advanced breeding lines and evaluation of virus level. Plant Disease, 81, 185-188.
 Go to original source...
Go to original source... - Lin, N.S., Hsu, Y.H. & Hsu, H.T. (1990) Immunological detection of plant viruses and a mycoplasmalike organism by direct tissue blotting on nitrocellulose membranes. Phytopathology, 80, 824-828.
 Go to original source...
Go to original source... - Lomonossoff, G.P. & Wege, C. (2018) TMV particles: the journey from fundamental studies to bionanotechnology applications. Advances in Virus Research, 102, 149-176.
 Go to original source...
Go to original source... - López-Moya, J.J., Fernández-Fernández, M.R., Cambra, M. & García, J.A. (2000) Biotechnological aspects of plum pox virus. Journal of Biotechnology, 76, 121-136.
 Go to original source...
Go to original source... - Luria, N., Smith, E., Sela, N. et al. (2018) A local strain of Paprika mild mottle virus breaks L3 resistance in peppers and is accelerated in Tomato brown rugose fruit virus-infected Tm-22-resistant tomatoes. Virus Genes, 54, 280-289.
 Go to original source...
Go to original source... - Maayan, Y., Pandaranayaka, E.P.J., Srivastava, D.A. et al. (2018) Using genomic analysis to identify tomato Tm-2 resistance-breaking mutations and their underlying evolutionary path in a new and emerging tobamovirus. Archives of Virology, 163, 1863-1875.
 Go to original source...
Go to original source... - Menzel, W. & Winter, S. (2021) Identification of novel and known tobamoviruses in tomato and other solanaceous crops using a new pair of generic primers and development of a specific RT-qPCR for ToBRFV. Acta Horticulturae, 1316, 143-148.
 Go to original source...
Go to original source... - Monroy-Borrego, A.G. & Steinmetz, N.F. (2022) Three methods for inoculation of viral vectors into plants. Frontiers in Plant Science, 13, 963756.
 Go to original source...
Go to original source... - Mulholland, V. (2009) Immunocapture-PCR for plant virus detection. In: Burns, R. (Ed.) Plant Pathology. Methods in Molecular Biology. Vol. 508. Totowa: Humana Press, pp. 183-192.
 Go to original source...
Go to original source... - Olmos, A., Capote, N., Bertolini, E. & Cambra, M. (2007) Molecular diagnostic methods for plant viruses. In: Punja, Z.K., de Boer, S.H., Sanfaçon, H. (Eds.) Biotechnology and Plant Disease Management. Wallingford: CABI Books, pp. 227-249.
 Go to original source...
Go to original source... - Plchová, H., Moravec, T., Čeřovská, N. et al. (2022) A GoldenBraid-compatible virus-based vector system for transient expression of heterologous proteins in plants. Viruses, 14, 1099.
 Go to original source...
Go to original source... - Plchova, H., Moravec, T., Hoffmeisterova, H., Folwarczna, J. & Cerovska, N. (2011) Expression of Human papillomavirus 16 E7ggg oncoprotein on N- and C-terminus of Potato virus X coat protein in bacterial and plant cells. Protein Expression and Purification, 77, 146-152.
 Go to original source...
Go to original source... - Rizzo, D., Da Lio, D., Panattoni, A. et al. (2021) Rapid and sensitive detection of tomato brown rugose fruit virus in tomato and pepper seeds by reverse transcription loop-mediated isothermal amplification assays (real time and visual) and comparison with RT-PCR end-point and RT-qPCR methods. Frontiers in Microbiology, 12, 640932.
 Go to original source...
Go to original source... - Salem, N., Mansour, A., Ciuffo, M., Falk, B.W. & Turina, M. (2016) A new tobamovirus infecting tomato crops in Jordan. Archives of Virology, 161, 503-506.
 Go to original source...
Go to original source... - Salem, N.M., Sulaiman, A., Samarah, N., Turina, M, & Vallino, M. (2022) Localization and mechanical transmission of tomato brown rugose fruit virus in tomato seeds. Plant Disease, 106, 275-281.
 Go to original source...
Go to original source... - Sarkes, A., Fu, H., Feindel, D., Harding, M. & Feng, J. (2020) Development and evaluation of a loop-mediated isothermal amplification (LAMP) assay for the detection of Tomato brown rugose fruit virus (ToBRFV). PLoS ONE, 15, e0230403.
 Go to original source...
Go to original source... - Sarrion-Perdigones, A., Falconi, E.E., Zandalinas, S.I. et al. (2011) GoldenBraid: an iterative cloning system for standardized assembly of reusable genetic modules. PLoS ONE, 6, e21622.
 Go to original source...
Go to original source... - Torrance, L. (1997) Developments in serological methods to detect and identify plant viruses. In: Cassells, A.C. (Ed.) Pathogen and Microbial Contamination Management in Micropropagation. Dordrecht: Springer, pp. 75-82.
 Go to original source...
Go to original source... - Towbin, H., Staehelin, T. & Gordon, J. (1979) Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets: procedure and some applications. Proceedings of the National Academy of Sciences of the United States of America, 76, 4350-4354.
 Go to original source...
Go to original source... - Vargas-Hernández, B.Y., Ramírez-Pool, J.A., Núñez-Muñoz, L.A. et al. (2022) Development of a droplet digital polymerase chain reaction (ddPCR) assay for the detection of Tomato brown rugose fruit virus (ToBRFV) in tomato and pepper seeds. Journal of Virological Methods, 302, 114466.
 Go to original source...
Go to original source... - Wang, Y., Dai, J., Liu, Y. et al. (2021) Development of a potential penside colorimetric LAMP assay using neutral red for detection of African swine fever virus. Frontiers in Microbiology, 12, 609821.
 Go to original source...
Go to original source... - Zaitlin, M. (1975) Tobacco mosaic virus (type strain). In: CMI/AAB Description of Plant Viruses. Kew: Commonwealth Mycological Institute.



 ORCID...
ORCID...